NOTE: We are working on migrating this site away from MediaWiki, so editing pages will be disabled for now.
JBrowse
From GMOD
Revision as of 20:49, 26 October 2011 by RobertBuels (Talk | contribs)
Status
- Mature release
- Active development
- Active support
Resources
JBrowse is a genome browser with an AJAX-based interface. JBrowse renders most tracks using client side JavaScript and JSON as its data transfer format. JBrowse is the official successor to GBrowse.
Contents
Demo
Requirements
Documentation
- JBrowseDev/Main - Thorough documentation for the most recent JBrowse release
- JBrowse Tutorial - Tutorial from 2009 GMOD Summer Schools
- JBrowse Home Page
- JBrowse Quick Tutorial - JBrowse administration.
Presentations
- Talk at the April 21, 2010 UCSC genome browser group ("genecats") meeting.
- JBrowse: A next-generation genome browser - Presentation by Ian Holmes at the August 2009 GMOD Meeting. See the talk's summary for more.
- JBrowse - Presentation by Mitch Skinner at the January 2009 GMOD Meeting. See the talk's summary for more.
- Presentation by Ian Holmes at July 2008 GMOD Meeting. See the talk's summary for more.
Downloads
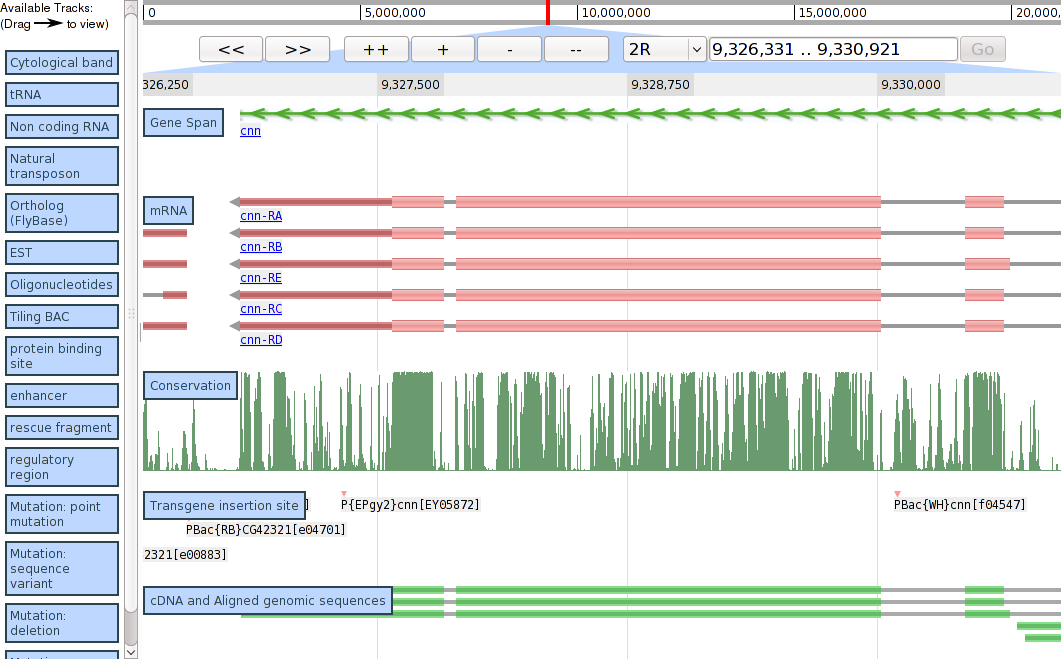
Screenshots
Demo screenshot from 2009/03/24, showing region of Drosophila melanogaster 2R.
JBrowse Development
JBrowse is an open-source project. Contributions are encouraged and welcomed. There is a developer mailing list and teleconferences on the 3rd Monday of the month at 2pm Pacific US time. Please contact Ian Holmes if you want to participate in the call.
Contact
- Mailing lists
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| JBrowse | gmod-ajax | JBrowse help and general questions. | Nabble (2010/05+), Sourceforge |
| jbrowse-dev | JBrowse development discussions. | Nabble (2011/08+) |
- Original author
- Mitch Skinner