Difference between revisions of "JBrowse/tool data"
RobertBuels (Talk | contribs) |
RobertBuels (Talk | contribs) |
||
| Line 59: | Line 59: | ||
3. Run the automated-setup script, <code>./setup.sh</code>, which will attempt to install all of JBrowse's (modest) prerequisites for you in the <code>jbrowse/</code> directory itself. Note that <code>setup.sh</code> does not need to be run as root or with <code>sudo</code>. | 3. Run the automated-setup script, <code>./setup.sh</code>, which will attempt to install all of JBrowse's (modest) prerequisites for you in the <code>jbrowse/</code> directory itself. Note that <code>setup.sh</code> does not need to be run as root or with <code>sudo</code>. | ||
| − | 4. Visit http:// | + | 4. Visit JBrowse on your machine, substituting the <nowiki>http://(your_machine/path_to_jbrowse)/index.html?data=sample_data/json/volvox</nowiki>. If you can see the included Volvox example data, you are ready to configure JBrowse to show your own data! |
| config = | | config = | ||
Revision as of 13:39, 29 April 2013
This page stores the data that populates the JBrowse wiki page.
{{ {{{template}}}
| name = JBrowse
| full_name =
| status = mature
| dev = active
| support = active
| os_linux = y
| os_mac = y
| os_unix = y
| os_win = y
| os_web = y
| logo = JBrowseLogo.png
| home = http://jbrowse.org
| about = JBrowse is a genome browser with a fully dynamic AJAX interface, being developed as the eventual successor to GBrowse. It is very fast and scales well to large datasets. JBrowse does almost all of its work directly in the user's web browser, with minimal requirements for the server.
Features
- Fast, smooth scrolling and zooming. Explore your genome with unparalleled speed.
- Scales easily to multi-gigabase genomes and deep-coverage sequencing.
- Supports GFF3, BED, FASTA, Wiggle, BigWig, BAM, and more.
- Very light server resource requirements. Serve huge datasets from a single low-cost cloud instance.
| public_server = | dl = | dl_url = http://jbrowse.org/install/ | dl_src = | dl_src_url = http://github.com/GMOD/jbrowse | dl_dev = | dl_dev_url = | getting_started_preamble = The JBrowse Quick-Start Tutorial provides a basic step-by-step recipe for quickly getting up and running with JBrowse. | req = JBrowse requires libpng, Zlib, and GD development libraries, plus make and a C compiler. On Ubuntu, you can install these prerequisites using the command:
sudo apt-get install libpng-dev libgd2-noxpm-dev build-essential
For tips on installing these baseline libraries, see JBrowse Troubleshooting.
| install = The JBrowse Quick-Start Tutorial provides a basic step-by-step recipe for quickly getting up and running with JBrowse.
1. Download JBrowse on to your web server.
2. Unpack JBrowse into a directory that is served by your web browser. On many systems, this defaults to /var/www.
cd /var/www unzip JBrowse-*.zip
Make sure you have permissions to write to the contents of the jbrowse/ directory you have just created.
3. Run the automated-setup script, ./setup.sh, which will attempt to install all of JBrowse's (modest) prerequisites for you in the jbrowse/ directory itself. Note that setup.sh does not need to be run as root or with sudo.
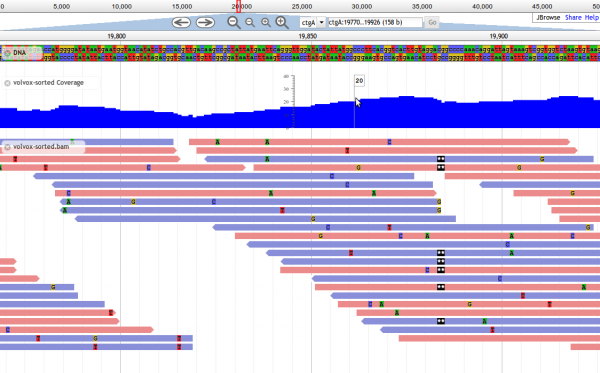
4. Visit JBrowse on your machine, substituting the http://(your_machine/path_to_jbrowse)/index.html?data=sample_data/json/volvox. If you can see the included Volvox example data, you are ready to configure JBrowse to show your own data!
| config = See the JBrowse Configuration Guide for information on:
- Formatting reference sequences (e.g. from FASTA files, or a Chado database)
- Feature Tracks (e.g. from BED or GFF files, a Chado database, or the UCSC genome browser)
- Image Tracks (e.g. from WIG files)
- Wiggle/BigWig Tracks
- Name Search and Autocompletion
- Removing tracks
- Compressing data stored on the server
- URL control
- Faceted track selection
- Anonymous usage statistics
Additional topics:
Upgrading JBrowse
To upgrade an existing JBrowse (1.3.0 or later) to the latest version, simply move its data directory (and jbrowse_conf.json if you are using it) into the directory of a newer JBrowse, and the new JBrowse will display that data.
To upgrade a 1.2.x JBrowse, copy its data directory into the new JBrowse directory, and point your browser at compat_121.html in the new JBrowse directory, instead of index.html.
If you are upgrading from a version of JBrowse older than 1.2.0, a fresh installation is required.
| doc = For data format specifications, developer information, and so forth, see JBrowse Advanced Topics. | papers = | presentations =
- April 2013 - Bio-IT World, Robert Buels: PDF
- August 2012 - presentation given as part of the 2012 GMOD Summer School: PDF
- April 2012 - GMOD 2012 Community Meeting, Robert Buels: PDF
- January 2012 - Plant and Animal Genome (PAG) XX: PDF
- April 2010 - UCSC genome browser group ("genecats") meeting: PDF
- August 2009 - GMOD Community Meeting: Talk summary | PDF
- January 2009 - GMOD Community Meeting: Talk summary | PDF
- July 2008 - GMOD Community Meeting: Talk summary | PDF
| tutorials =
- Getting Started with JBrowse Tutorial
- part of the JBrowse documentation
| wild_urls = | mail = Please direct questions and inquiries regarding JBrowse to the mailing lists below.
Requests for help should be directed to gmod-ajax@lists.sourceforge.net.
| Mailing List Link | Description | Archive(s) | |
|---|---|---|---|
| JBrowse | gmod-ajax | JBrowse help and general questions. | Nabble (2010/05+), Sourceforge |
| jbrowse-dev | JBrowse development discussions. | Nabble (2011/08+) |
| logo_info = | dev_ppl = JBrowse is an open-source project, started and managed by the laboratory of Ian Holmes at the University of California, Berkeley.
As of January 2012, the lead developer of JBrowse is Robert Buels. Most of JBrowse was originally written by Mitch Skinner.
There is a mailing list for developers, and there is usually a teleconference on the 3rd Monday of the month at 2pm Pacific US time. We welcome participation from anyone and everyone. Please contact Robert Buels if you would like to listen in or participate.
| dev_status = The JBrowse source code repository is kept on GitHub. Please feel very free to fork the code on GitHub and make modifications and improvements, submitting pull requests. GitHub has a very nice tutorial on how to get started with this style of development.
| contact_email = rbuels@gmail.com | input = GFF3, BED, FASTA, Wiggle, BigWig, BAM | output = | see_also = | demo_server = Volvox example data | extras =
Taught at the 2013 GMOD Summer School
Included in
}}