GMOD
Debian Stable Installation Notes
Contents
Debian Stable (Etch 4.0) installation notes
Installing Chado/GMOD old office machine with a fresh Debian stable installation. Goal: test the Chado scheme for annotation of bacterial genomes
- AMD Athlon(tm) XP 3200+
- 1 GB memory
- 500 GB SATA disk
I borrowed a lot of stuff from the Installing_Chado_on_Ubuntu_HOWTO page as Debian and Ubuntu are much alike.
Apache
Apache 2.2 is installed by default during the installation of Debian stable (web package).
location for GMOD
mkdir /usr/local/gmod
note: this is NOT the home of my gmod user.
download
got the source from sourceforge and unpacked in /usr/local/gmod (maybe I should use the cvs version??)
=== postgresql setup ===
Note: Debian stable has postgresql 7.4 as default package, but i’m going for 8.1.11 as this is also the version that runs on our RedHat Enterprise 5 server (more on that later)
apt-get install the following packages, Synaptic is your friend
- postgresql-8.1
- postgresql-8.1-client-common
- postgresql-client-8.1
- postgresql-contrib-8.1
- postgresql-plperl-8.1
Not necessary, but I also installed phpPgAdmin from source, not the Debian package.
Following the guidelines from INSTALL.Chado to add a database user and optimize some settings. Don’t forget to create a database user root as well for the CPAN installations below, otherwise some tests fail. I added user ‘chado’ for the chado database installation.
The configuration files are located in a different location as opposed to our Redhat installation:
/etc/postgresql/8.1/main
I opted to install the CPAN modules instead of the Debian packages as some of the Debian packages have older versions of the modules. The modules are installed in
/usr/local/share/perl/5.8.8/
So not overwriting any installed Debian packages.
Note: when installing DBIx::DBStag, the install tests require a postgres account to run the ‘make tests’. Because CPAN installs are typically run under root, you may create an account for root with:
su postgres createuser root
Superuser[Y]
The DBIx::DBStag package should now be able to install without using the ‘force’ option under CPAN. For security, don’t forget to drop the user after the install is finished:
su postgres dropuser root
CPAN packages
CGI
GD
DBI
DBD::Pg
SQL::Translator
Digest::MD5
Text::Shellwords
Graph
Data::Stag
XML::Parser::PerlSAX
Module::Build
Class::DBI
Class::DBI::Pg
Class::DBI::Pager
DBIx::DBStag (Force install needed if tests fail)
XML::Simple
LWP
Template
Log::Log4perl
GO::Parser
BioPerl
I installed BioPerl-live using subversion into /usr/local/bioperl/bioperl-live (as root)
svn co svn://code.open-bio.org/bioperl/bioperl-live/trunk bioperl-live
cd /usrlocal/bioperl/bioperl-live
perl Build.PL
./Build
./Build test #some tests fail as usual
sudo ./Build install
Environment Variables
Added this to my .bashrc so that when I login as user gmod all vars are set
# GMOD additions
export GMOD_ROOT=/usr/local/gmod
export GO_ROOT=/usr/local/share/perl/5.8.8/GO/
export PERL5LIB=$PERL5LIB:/usr/local/bioperl/bioperl-live
export CHADO_DB_NAME=dev_chado_01
export CHADO_DB_USERNAME=chado
export CHADO_DB_PASSWORD=******
#CHADO_DB_HOST=localhost
#CHADO_DB_PORT=5432
=== installing Chado scheme and tools
in /usr/local/gmod
perl Makefile.PL
make
sudo make install
make load_schema
make prepdb
a quick look at the database user phpPgAdmin: the database looks fine.
make ontologies
./Build ontologies
Available ontologies:
[1] Relationship Ontology
[2] Sequence Ontology
[3] Gene Ontology
[4] Chado Feature Properties
[5] Cell Ontology
[6] Plant Ontology
Which ontologies would you like to load (Comma delimited)? [0] 1,2,3,4
This take a while. i’m getting the notorious duplicate key violates unique constraint “cvterm_c1” error as well.
Data loading
We have a local mirror of [1] mounted at /data/genomes As described in the Chado manual I inserted an organism in the database, our pet organism of course:
INSERT INTO organism (abbreviation, genus, species, common_name)
VALUES ('L.plantarum', 'Lactobacillus', 'plantarum', 'Lactobacillus plantarum WCFS1');
question: other pages seem to specify the taxonomy id as organism id. In our case that would be 220668. Should we use this ID or just let postgresql add an ID for us?
what doesn’t work
NCBI gff3 files (does not work!)
> gmod_gff3_preprocessor.pl -gfffile /data/genomes/Lactobacillus_plantarum/NC_004567.gff -outfile NC_004567.gff
fails with
"my" variable %seen masks earlier declaration in same scope at /usr/local/share/perl/5.8.8/Bio/GMOD/DB/Adapter.pm line 4199.
"my" variable %seen masks earlier declaration in same scope at /usr/local/share/perl/5.8.8/Bio/GMOD/DB/Adapter.pm line 4223.
NOTICE: CREATE TABLE will create implicit sequence "gff_sort_tmp_row_id_seq" for serial column "gff_sort_tmp.row_id"
NOTICE: CREATE TABLE / PRIMARY KEY will create implicit index "gff_sort_tmp_pkey" for table "gff_sort_tmp"
couldn't open /data/genomes/Lactobacillus_plantarum/NC_004567.gff.sorted.fasta for writing: Read-only file system
although running from a writable directory, gmod_gff3_preprocessor.pl tries to write to the location where the input file is.
> cp /data/genomes/Lactobacillus_plantarum/NC_004567.gff NC_004567.gff
> gmod_gff3_preprocessor.pl --gfffile NC_004567.gff --outfile NC_004567.gff.sorted
"my" variable %seen masks earlier declaration in same scope at /usr/local/share/perl/5.8.8/Bio/GMOD/DB/Adapter.pm line 4199.
"my" variable %seen masks earlier declaration in same scope at /usr/local/share/perl/5.8.8/Bio/GMOD/DB/Adapter.pm line 4223.
Sorting the contents of NC_004567.gff ...
Writing sorted contents to NC_004567.gff.sorted
>gmod_bulk_load_gff3.pl -organism 'Lactobacillus plantarum WCFS1' --gfffile NC_004567.gff.sorted
This results in lots of errors
(Re)creating the uniquename cache in the database...
Creating table...
Populating table...
Creating indexes...Done.
Preparing data for inserting into the dev_chado_01 database
(This may take a while ...)
There is a CDS feature with no parent (ID:) I think that is wrong!
This GFF file has CDS and/or UTR features that do not belong to a
'central dogma' gene (ie, gene/transcript/CDS). The features of
this type are being stored in the database as is.
Unable to find srcfeature NC_004567.1 in the database.
Perhaps you need to rerun your data load with the '--recreate_cache' option. at /usr/local/share/perl/5.8.8/Bio/GMOD/DB/Adapter.pm line 4026
Bio%253A%253AGMOD::DB::Adapter::src_second_chance('Bio%253A%253AGMOD::DB::Adapter=HASH(0x8bba494)', 'Bio::SeqFeature::Annotated=HASH(0x8c4be6c)') called
at /usr/local/bin/gmod_bulk_load_gff3.pl line 758
Issuing rollback() for database handle being DESTROY'd without explicit disconnect().
Loading NCBI gff files directly is not going to work.
NCBI genbank RefSeq files (does work, with adaptations) Please note that the page at Load_RefSeq_Into_Chado is now including an organism_id, confusing.
taken from RefSeq howto –> does not load in the database
bp_genbank2gff3.pl --summary --typesource chromosome -noCDS --filter exon -o . NC_004567.gbk
What works
taken from GenBank howto –> does load in the database
bp_genbank2gff3.pl -noCDS -s -o . NC_004567.gbk
Gives some output
# Input: NC_004567.gbk
# working on region:NC_004567, Lactobacillus plantarum WCFS1, 03-DEC-2007, Lactobacillus plantarum WCFS1, complete genome.
# GFF3 saved to ./NC_004567.gbk.gff
# Summary:
# Feature Count
# ------- -----
# mRNA 3007
# gene 3093
# region 1
# pseudogene 42
# CDS 3007
# RESIDUES(tr) 910927
# RESIDUES 3308274
# rRNA 16
# pseudogenic_region 42
# exon 3093
# tRNA 70
Load into the database with:
gmod_bulk_load_gff3.pl --organism 'Lactobacillus plantarum WCFS1' --dbxref GeneID --gfffile NC_004567.gbk.gff --recreate_cache
load the file into the database–>ok
Note: one could use the following options for the bp_genbank2gff3.pl script: –typesource chromosome –typesource plasmid etc, as long as it is a valid SO type.
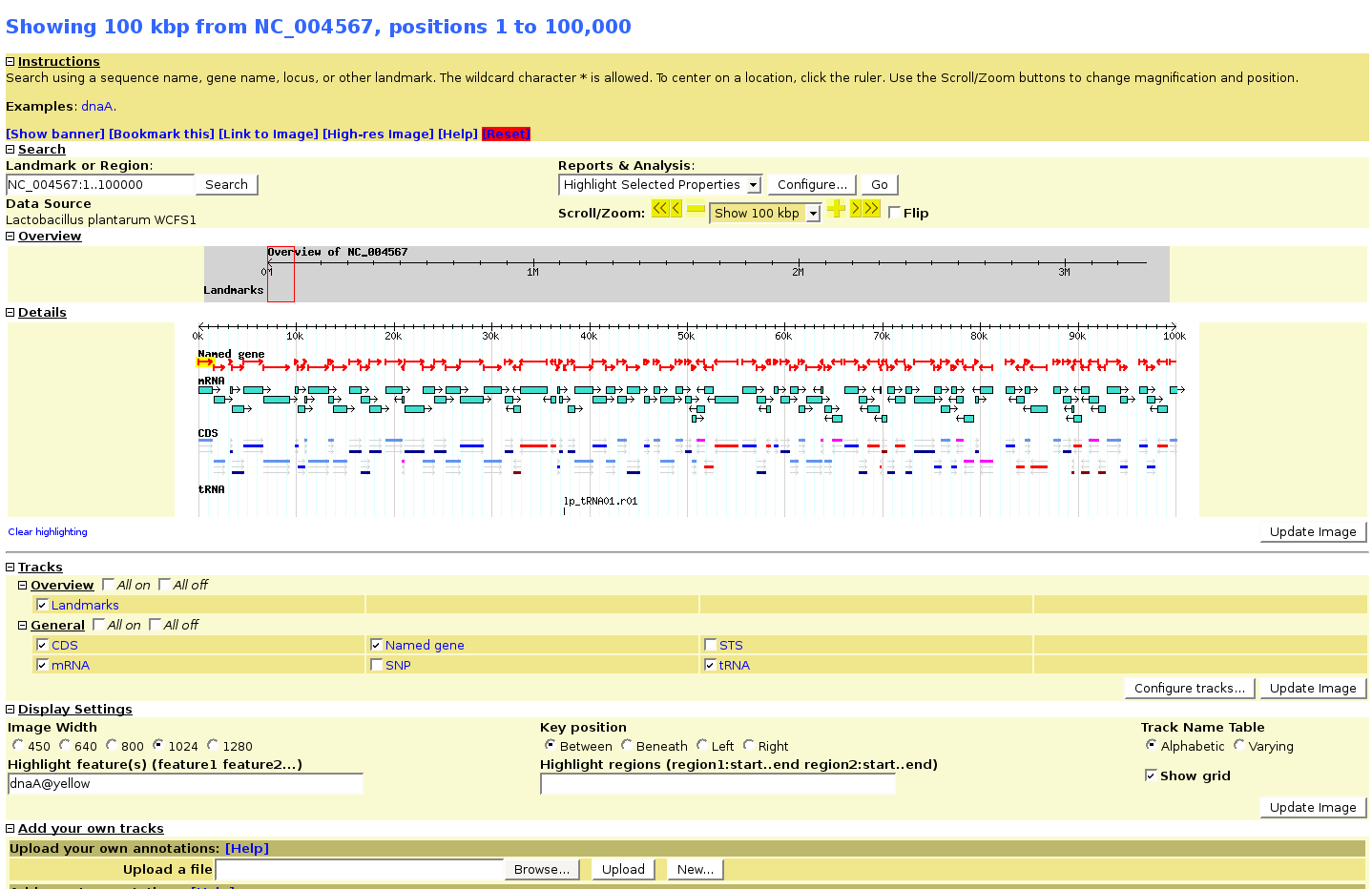
Gbrowse configuration
completely standard as described in the manual. Configuration file goes in /etc/apache2/