GMOD
Tripal Tutorial 2010

This Tripal tutorial was presented by Stephen Ficklin at the 2010 GMOD Summer School - Americas, May 2010. The most recent Tripal tutorial can be found at the Tripal Tutorial page.
This tutorial walks you through installing and configuring Tripal, a web front end to Chado databases. This tutorial references (and complements) the Tripal User’s Guide, January 2010 edition.
Contents
- 1 VMware
- 2 Caveats
- 3 Important Information
- 4 Pre-Course Setup
- 5 Software Used
- 6 Pre-Start Tasks
- 7 Install Drupal
- 8 Re-enable GBrowse, JBrowse, …
- 9 Review the features of Drupal
- 10 Setup Drupal Cron
- 11 Theming
- 12 Tripal Resources
- 13 Tripal Overview
- 14 Setup Tripal
- 15 Organisms
- 16 Features
- 17 Analysis Modules
- 18 GO Module
- 19 Blast Module
- 20 InterPro Analysis
- 21 Searching
- 22 Libraries & other Analysis
- 23 Incorporate JBrowse
- 24 Example Customizing Content
- 25 Writing your own Module
VMware
This tutorial was taught using a VMware system image as a starting point. If you want to start with the same system, download and install the start image (below). See VMware for what software you need to use a VMware system image and for directions on how to get the image up and running on your machine.
Download the start image and the end image.
Logins:
| Purpose | Username | Password |
|---|---|---|
| Shell | gmod | gmodamericas2010 |
| MySQL | root | gmodamericas2010 |
Caveats
Important Note
This tutorial describes the world as it existed on the day the tutorial was given. Please be aware that things like CPAN modules, Java libraries, and Linux packages change over time, and that the instructions in the tutorial will slowly drift over time. Newer versions of tutorials will be posted as they become available.
Important Information
- Tripal is open source and currently at version 0.2 (second release).
- The documentation is complete enough to allow for setup of a site.
- The API for adding customized modules will need to be added to the documentation.
- A publication for Tripal is currently in the works and should be submitted this summer.
- Data loaded into chado for this course was taken from the FlyBase website
- It’s new software. There are still bugs…
Pre-Course Setup
By way of information the following steps were performed prior to the course. These steps will not be needed during the class but are here for reference.
Software Used
- Apache Web server
- PHP (both command-line and web module)
- PostgreSQL database management system
- phpPgAdmin (for easy database administration)
- Chado
- BioPerl
- go-perl
- Drupal
- Tripal
Web Prep
Install & Setup PHP
sudo apt-get install php5
sudo apt-get install php5-pgsql
sudo apt-get install php5-cli
sudo apt-get install php5-gd
Change some php settings (as root):
cd /etc/php5/apache2
sudo gedit php.ini
Set the memory_limit to something larger than 16M (should not exceed
physical memory, be conservative but not too much so):
memory_limit = 2048M;
Now, restart the webserver:
sudo /etc/init.d/apache2 restart
Do the same for the command-line php.ini:
cd /etc/php5/cli/
sudo gedit php.ini
Set the memory limit:
memory_limit = 2048M;
Install phpPgAdmin
phpPgAdmin is a nice web-based utility for easy administration of a PostgreSQL database. It is not required for successful operation of Tripal but is very useful.
cd /home/gmod/Documents/Software/tripal/packages
wget http://downloads.sourceforge.net/phppgadmin/phpPgAdmin-4.2.3.tar.gz?download
As root:
cd /var/www
sudo tar -zxvf /home/gmod/Documents/Software/tripal/packages/phpPgAdmin-4.2.3.tar.gz
sudo ln -s phpPgAdmin-4.2.3/ phppgadmin
Copy the conf/config.inc.php-dist to conf/config.inc.php
cd phppgadmin/conf
sudo cp config.inc.php-dist config.inc.php
Set permission for the web user:
cd /var/www
sudo chgrp -Rh www-data phppgadmin
sudo chgrp -Rh www-data phpPgAdmin-4.2.3/
Prepare Postgres
We need to create two Postgres databases, one for the Chado tables and the other for the Drupal tables.
sudo su - postgres
Create the user that will manage the database:
postgres@gmod:~$ createuser -P fly_admin
Enter password for new role: flydemo2010
Enter it again: flydemo2010
Shall the new role be a superuser? (y/n) n
Shall the new role be allowed to create databases? (y/n) y
Shall the new role be allowed to create more new roles? (y/n) n
Create the chado and drupal databases:
postgres@gmod:~$ createdb chado_dmel -O fly_admin
postgres@gmod:~$ createdb drupal_dmel -O fly_admin
Install Prereqs
XSLT
sudo apt-get install xsltproc
Install the Postgres development tools:
sudo apt-get install libpq-dev
Install the needed perl modules
sudo perl -MCPAN -e shell
install GO::Parser
install Template
install XML::Simple
install Log::Log4perl
install XML::Parser::PerlSAX
install DBI
install DBD::Pg
install DBIx::DBSchema
install DBIx::DBStag
install Parse::RecDescent
Needed for CUGI scripts (optional, use of CUGI scripts is not needed for this demonstration)
install Spreadsheet::WriteExcel
Download and install BioPerl:
cd /home/gmod/Documents/Software/tripal/packages/
wget http://bioperl.org/DIST/BioPerl-1.6.1.tar.gz
tar -zxvf BioPerl-1.6.1.tar.gz
cd BioPerl-1.6.1/
perl Makefile.PL
make
sudo make install
Install the go-perl modules:
cd /home/gmod/Documents/Software/tripal/packages/
wget http://search.cpan.org/CPAN/authors/id/C/CM/CMUNGALL/go-perl-0.10.tar.gz
tar -zxvf go-perl-0.10.tar.gz
cd go-perl-0.10
perl Makefile.PL
make
sudo make install
Chado Installation
Download and extract the gmod package:
cd /home/gmod/Documents/Software/tripal/packages/
wget http://internap.dl.sourceforge.net/sourceforge/gmod/gmod-1.0.tar.gz
tar -zxvf gmod-1.0.tar.gz
cd gmod-1.0
Set some environment variables that provide database connection for Chado:
export GMOD_ROOT=/home/gmod/Documents/Software/tripal/gmod-1.0
export CHADO_DB_NAME=chado_dmel
export CHADO_DB_USERNAME=fly_admin
export CHADO_DB_PASSWORD=flydemo2010
export CHADO_DB_HOST=localhost
export CHADO_DB_PORT=5432
Now install the gmod package and pre-populate the database:
perl Makefile.PL PREFIX=/home/gmod/Documents/Software/tripal/gmod-1.0
make
make install
make load_schema
make prepdb
make ontologies
Load DMel Data
Download the GFF file available (originally) from the fly genome website. For the sake of simplicity we will only be using chromosome 4:
cd /home/gmod/Documents/Data/tripal
wget ftp://ftp.gmod.org/pub/gmod/Courses/2010/SummerSchoolAmericas/Tripal/dmel-4-r5.27.gff.gz
Download the coding segments:
wget ftp://ftp.gmod.org/pub/gmod/Courses/2010/SummerSchoolAmericas/Tripal/dmel-4-CDS-r5.27.fasta.gz
Adjust the
/home/gmod/Documents/Software/tripal/gmod-1.0/bin/gmod_bulk_load_gff3.pl
so it can find the gmod libraries (we didn’t install in a default
location):
use lib "/home/gmod/Documents/Software/tripal/gmod-1.0/share/perl/5.10.0";
A filtered version of the downloaded GFF file was preloaded into the database for the course. However, the data was not loaded how we wanted. See the section below for removing these features and reloading.
Prepare Apache
Enable the rewrite module for apache. This is useful so that we can use Clean URLs with Drupal. Clean URLs are not required but make the page URLs easier to use:
cd /etc/apache2/mods-enabled
sudo ln -s ../mods-available/rewrite.load
Pre-Start Tasks
Reload Fly Data
We need to reload the fly data that was loaded previous to class. The GFF file was not quite compatible with how we want to display the data.
First log on to Postgres and remove all features from the database
First delete all features from Chado:
psql -U fly_admin -d chado_dmel -q -c "delete from feature"
Next, download a GFF file that has been reduced and adjusted for the course:
Media:Dmel-4-r5.27.reduced.gff.txt
The following changes were made from the original GFF:
- The order of features in the GFF confused the Chado bulk loader so the features were rearranged.
- Only features of type ‘gene’, ‘pseudogene’, ‘CDS’,’intron’ and ‘mRNA’ are included.
- Feature IDs and Feature Names were set as the same (previously they were not the same and caused the bulk load script to create two genes with the same name which confused Drupal).
- Gene annotations were removed since they were imported into chado as feature synonyms. Tripal does not consider annotations as synonyms and we will display annotations through analyses.
Rename the file take off the .txt extension and
cd /home/gmod/Downloads
mv Dmel-4-r5.27.reduced.gff.txt Dmel-4-r5.27.reduced.gff
Finally, load the new data:
export GMOD_ROOT=/home/gmod/Documents/Software/tripal/gmod-1.0
$GMOD_ROOT/bin/gmod_bulk_load_gff3.pl \
--dbname chado_dmel \
--dbuser fly_admin \
--dbhost localhost \
--dbpass flydemo2010 \
--organism fruitfly \
--gff Dmel-4-r5.27.reduced.gff \
--recreate_cache \
--no_target_syn
Prepare for GO Terms
This step would normally not occur at this point but it takes about 20 minutes to prepare the database for GO terms. Tripal has an interface to do this but to save time and keep the course moving we will do this manually now:
psql -U fly_admin -d chado_dmel -q -c "SELECT * FROM fill_cvtermpath('biological_process')" &
psql -U fly_admin -d chado_dmel -q -c "SELECT * FROM fill_cvtermpath('cellular_component')" &
psql -U fly_admin -d chado_dmel -q -c "SELECT * FROM fill_cvtermpath('molecular_function')" &
check to make sure these are running:
jobs
Now open a new terminal and leave this alone.
Install Drupal
cd /var/www
sudo mv index.html index.old.html
sudo tar -zxvf /home/gmod/Documents/Software/tripal/drupal-6.16.tar.gz
Now that we’ve unpacked Drupal, we’ll move some things around.
cd drupal-6.16
sudo mv .htaccess * ../
Set permissions so we can easily work with
cd /var
sudo chown -R gmod:www-data www
This will undo the permission changes we did previously with GBrowse. Redo them.
sudo chmod -R 777 /var/www/gbrowse2/{tmp,databases}
and maybe other dirs as well?
Be more careful in real life.
Configure Drupal
cd /var/www/sites/default
cp default.settings.php settings.php
Now edit the settings.php file using your favorite editor (e.g. vi,
gedit, emacs).
Change the $db_url argument to be:
$db_url = array (
'default' => 'pgsql://fly_admin:flydemo2010@localhost/drupal_dmel',
'chado' => 'pgsql://fly_admin:flydemo2010@localhost/chado_dmel'
);
Create a files directory and make it writable by the webserver
mkdir files
sudo chgrp www-data files
chmod g+rw files
Edit the configuration file and change the AllowOverride from None
to All.
cd /etc/apache2/sites-enabled
sudo vi 000-default
<Directory /var/www/>
Options Indexes FollowSymLinks MultiViews
AllowOverride All
Order allow,deny
allow from all
</Directory>
Restart the web server
sudo /etc/init.d/apache2 restart
Point Browser to the webserver and follow the two easy pages for installing and then configuration the Drupal site.
http://localhost/install.php
The username and password used in this course were
- Username: fly_admin
- Password: flydemo2010
Re-enable GBrowse, JBrowse, …
At this point, the GBrowse, JBrowse, and other web pages that were set up earlier, no longer work. Drupal broke them. Fix them.
sudo gedit /var/www/.htaccess
Change
# Set the default handler.
DirectoryIndex index.php
to
# Set the default handler.
DirectoryIndex index.php index.html
Review the features of Drupal
User Accounts
- View History
- Edit the Account
Create Content
- Create a ‘Home’ page
- Create an ‘About’ page
Administration - Content Management
- Content Management → Content
- Content Management → Taxonomy
Administration - Site Building
- Site Building → Blocks
- Add the “Who’s Online” block to the left sidebar.
- Add a search box to the left sidebar as well.
- Site Building → Menu
- Add a ‘Home’ link
- Add an ‘About’ link
- Site Building → Modules
- Tripal requires the Path and Search modules. Enable those.
- Return to the About link and add an alias.
- Return to the Menu and update the about link.
- Navigate to Drupal.org and click on ‘Modules’. Review the wealth of public modules.
- Site Building → Themes
- Change the theme to Pushbutton.
- Configure the theme and remove the search box.
- Navigate to Drupal.org and click on ‘Themes’. Review the wealth of public themes.
Administration - User Management
- User Management → Roles
- Create role ‘webmaster’
- Create role ‘editor’
- User Management → Users
- Add a user for yourself and set as webmaster.
- User Management → Permissions
- Turn on everything for webmaster
- Set ‘Access Content’ for editor (and whatever else)
- User Management → User Settings
Administration - Site Configuration
- Site Configuration → Performance
Administration - Reports
- Reports → Recent log entries
- Reports → Status reports
- Run cron manually
Setup Drupal Cron
The Drupal cron is used to automatically execute necessary housekeeping tasks on a regular interval.
Drupal requires an entry in the crontab to function:
crontab -e
A word on text editors such as nano.
Add this line to the crontab
0,30 * * * * /usr/bin/wget -O - -q http://localhost/cron.php > /dev/null
The cron will launch this job every 30 minutes.
Theming
Drupal allows for customization for the look-and-feel of the website. This is done through use of themes. A list of publicly available themes is available on the Drupal website.
Themes not packaged with Drupal should be installed in a special
directory. Create the themes directory:
cd /var/www/sites/all
mkdir themes
cd themes
For this demonstration we will use the Pixture Reloaded theme
(http://drupal.org/node/456520). Installation is easy.
Simply download and then unpack the downloaded compressed theme file
into the newly created themes directory:
wget http://ftp.drupal.org/files/projects/pixture_reloaded-6.x-3.2.tar.gz
tar -zxvf pixture_reloaded-6.x-3.2.tar.gz
Now, to enable this theme, navigate to the Administer → Site Building → Themes page and select the Pixture Reloaded radio button and checkbox, remove the check on the ‘Garland’ theme (default theme) and click Save configuration.
This theme allows you to change the colors. Click the configure link next to the theme and change the color scheme.
As an example of customizability of themes let’s change the logo image. Download this image and upload as the logo:

Tripal Resources
Availability
- Current package available for download from CUGI website.
- Active source code available for download from Subversion at the GMOD Sourceforge site.
Documentation and Helper Scripts
Documentation and helper scripts can be obtained on the CUGI website.
Mailing List
Made available through GMOD: https://lists.sourceforge.net/lists/listinfo/gmod-tripal
Tripal Overview
Module Overview
The following image shows a rough overview of the hierarchy of modules in Tripal. Not all dependency relationships are shown but the order shows the general organizational structure of the modules. Also, modules in Tripal are divided into three classes: a core module, chado-centric modules and analysis modules
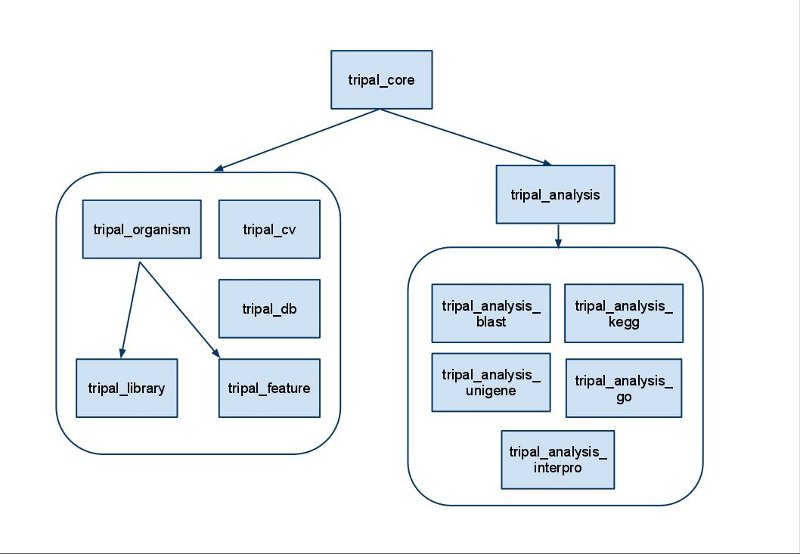
Tripal Core Module
The core module provides the following functionality:
- Jobs – long running items
- Taxonomy – for advanced searching
- CVterms – for customized data
- Materialized Views – for speedy
- API for new modules – for expandability
Chado-centric Modules
Chado-centric modules provide “wrappers” for chado modules:
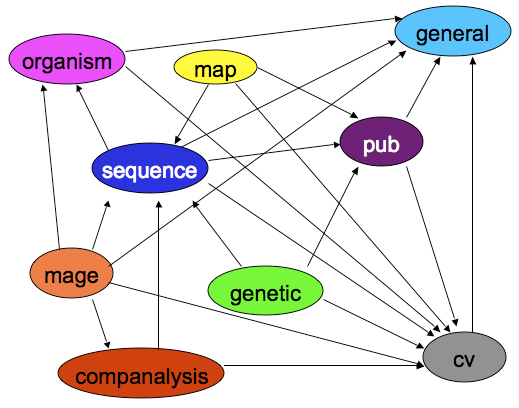
Analysis Modules
Analysis modules are intended to be the most diverse and customized for each site’s needs. Currently, CUGI has written several analysis modules that we have used for development of sites we manage. These include:
- Blast – manages storage and display of blast results
- Unigene – manages storage of features from a transcriptome unigene assembly.
- KEGG – manages storage and display of KEGG results for features as well as browseable trees for analyses.
- InterProScan – manage storage and dispaly of InterProScan results for features as well contribute GO terms
- Gene Ontology (GO) – provides trees and charts for visualization of GO term assignments as well as listing assigned GO terms on feature pages.
We highly encourage others to alter these as needed or write their own for custom analyses. Custom developed modules can be shared with the community.
Setup Tripal
Install Tripal
Create the necessary directory structure
cd /var/www/sites/all
mkdir modules
cd modules
Unpack the Tripal package into the modules directory:
tar -zxvf /home/gmod/Documents/Software/tripal/packages/tripal-6.x-0.2.tar.gz
Drupal wants to find all 3rd party modules in the modules directory so
we need to move our modules out of the tripal directory that we just
unpacked:
mv tripal-6.x-0.2/* ./
Tripal includes a “base” theme for Drupal to allow for customization of
the look-and-feel of
Chado data. This
base theme is provided in the theme_tripal directory. Move the theme
to the themes directory created earlier:
mv theme_tripal ../themes/
cd ../themes
We will enable the Tripal theming a bit later.
Enable the Tripal Modules
Navigate to the Modules Administration Page (Administer → Site Building → Modules) and click the checkboxes for each of the Tripal modules. Do not enable them all at once. Enable in groups:
Group 1 (dependencies, already enabled):
Path
Search
Group 2:
Tripal Core
Group 3:
Tripal Chado CV
Tripal Chado DB
Tripal Chado Organism
Tripal Chado Feature
Tripal Chado Library
Tripal Chado Analysis
Setup Tripal Cron
Tripal also require an entry in the crontab to function:
crontab -e
A word on text editors such as nano.
Add this line to the crontab
0,15,30,45 * * * * (cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin ) > /dev/null
This will run the Tripal cron every 15 miutes.
Test the cron job to make sure it works:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
The Tripal job should execute and return the following:
Tripal Job Launcher
-------------------
Calling: tripal_update_mview(3, 1)
One of the modules that we loaded previously uses materialized views
which we will discuss in a bit. The module added a job to the jobs
management system. The message ‘Calling: tripal_update_mview’ shows
that a job was executed.
Note: It is important the the final parameter be set to the name of
the administrator account for Drupal. In our case ‘fly_admin’
Organisms
Sync the Organism
What is syncing? Syncing is needed to create pages in Drupal that correspond to data (e.g. organisms and features) in Chado.
Navigate to Administer → Tripal Management → Organisms. Click the checkbox beside Drosophila melanogaster (fruitfly) and then click Sync Organisms. Afterwards, click on the link to go to the jobs page and view the job that has been submitted.
Rather than wait for at most 15 minutes for the cron to launch the job, we’ll manually perform that now:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Refresh the jobs page and notice the stats.
Edit the Organism
Now, the organism has a Drupal page! We can view it, but notice it has no useful content.. we need to add some. Click the Edit tab at the top. For a description add the following (stolen from wikipedia):
The genome of D. melanogaster (sequenced in 2000, and curated at the FlyBase database) contains four pairs of chromosomes: an X/Y pair, and three autosomes labeled 2, 3, and 4. The fourth chromosome is so tiny that it is often ignored, aside from its important eyeless gene. The D. melanogaster sequenced genome of 165 million base pairs has been annotated and contains approximately 13,767 protein-coding genes which comprise ~20% of the genome out of a total of an estimated 14,000 genes. More than 60% of the genome appears to be functional non-protein-coding DNA involved in gene expression control. Determination of sex in Drosophila occurs by the ratio of X chromosomes to autosomes, not because of the presence of a Y chromosome as in human sex determination. Although the Y chromosome is entirely heterochromatic, it contains at least 16 genes, many of which are thought to have male-related functions.
Also, download the image below and use the image upload field to include this image
Now, save the organism.
But, our additional information did not show up! We need to install the Tripal theme.
Setup the Tripal Theme
Enable the Base Theme
The Tripal theme is a base theme which adds functionality for Chado data
to the selected theme. To enable the Tripal theme we must edit the
configuration file for the Pixture Reladed theme. To do this, edit the
pixture_realoaded.info file that is included with the theme:
cd /var/www/sites/all/themes/pixture_reloaded
Now add the following to the pixture_reloaded.info
base theme = tripal
Now, navigate to the Administer → Site Building → Themes directory and enable the Tripal theme but do not set as default.
Return to the organism page and refresh. Now the content is available.
Organize the Blocks
The next step is to place the blocks in their proper place. Open a new tab in the browser and navigate to the Administer → Site Building → Blocks page. The theme has now changed to the admire-gray theme, but with Tripal “extensions”.
Alter the blocks accordingly:
Powered by Drupal → <none>
Libraries → Sidebar left
Organisms → Sidebar left
Search form → Sidebar left
Now, move the Search form and the Organism’ block to the top of the list of the Sidebar left group by clicking and dragging the crossbars. This will ensure our organism block shows up on the top of the list.
Click Save blocks. Now we have some blocks along the left-hand side of the site for easy access to Chado data.
Features
Sync the Features
- Navigate to the Administer → Tripal Management → Features. Before syncing any features we must first set the accession prefix and the feature types. Each feature on the site is assigned a unique accession number. Essentially this is the feature_id of the feature in the chado table. The prefix is added to the beginning of the feature_id to form the unique accession. The default is ‘ID’. Let’s change this to DMEL.
- Next, the site will only create drupal content for those features
types listed in the Feature Types text box. Replace the contents
of this box with
geneandpsuedogene. Then click the Sync Features button further down the page. Click on the link to go to the jobs page and view the job that has been submitted.
- Finally, we need to sync the features, click the Sync all Features button. Rather than wait on the job cron we’ll launch this job manually:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Note This can take a long time for large datasets. However for this demo our dataset has been reduced and it runs rather quickly. Check the status of the sync by navigating to the Administer → Tripal Management → Jobs page. It took about less than 1 minute on my laptop.
The features are synced but we have no way to find them. The administer can see them by navigating to Administer → Content Management → Content. However, we need a way for users to find features. There are two ways to do this:
- Add a browseable list of features (good for small sites)
- Searching
Feature Materialized Views
The feature module is capable of showing the type and number of features for this organism. This display is added to the organism page. Even after syncing of organisms and features this information does not appear on the Organism page. This behavior is controlled by a materialized view that is created when the module is installed but needs to be updated.
To view this materialized view, navigate to the Administer → Tripal Management → Materialized Views page. Click view on the organism_feature_count materialized view.
To update this view, click the update link at the bottom. Normally we would wait for the cron job. Rather than wait for the cron to launch the job, do it manually now:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Navigate to the Administer → Tripal Management → Features page and click the button Set Summary in the Feature Summary section.
Return to the organism page and notice that a new box appeared with a listing of the types and number of features for the organism as well as a pie chart.
Browse Features
Another potentially useful feature is a browseable list of features in the database. This also appears on the organism page, but controlled by the feature module.
To show the list of features, navigate to the Administer → Tripal Management → Features. Click the button Set Browser in the Feature Browser section.
Note: This browseable list can cause organism pages to load slowly for large sites. This is a problem with the way Drupal handles paged lists.
Enable Searching of Features
Aside from the browseable list we cannot locate features for viewing without searching. To index features for searching, navigate to Administer → Tripal Management → Features. Click on the button Reindex all feature nodes.
Rather than wait on the Drupal cron to execute our job we’ll run it manually:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Now, search for the feature ‘FBgn0040037’.
If desired, features can be edited. Useful for small fixes, but not good for bulk data changes.
Feature References
Notice on the bottom of the page a References box exists. We would like for this to have a link to the external FlyBase database. To do this navigate to Administer → Tripal Management → DB. The page with ‘FlyBase’ is on page 5. Edit and add the following:
# URL: http://flybase.org/
# URL prefix: http://flybase.org/reports/
Now return to the feature page and the reference should be clickable
Review Permissions
Now that we have data in the site we can control who has access to do certain things. Navigate to Administer → User Management → Permissions. Review the available permissions for anonymous users, webmaster and editor.
Analysis Modules
GO Module
The GO module provides charts and trees for browsing GO assignments. It also lists GO assignments on the feature pages.
At this we can would normally prepare the database to handle GO terms. This takes several tens of minutes so we did this manually at the start of the course. But to see how this would normally be done, navigate to Administer → Tripal Management → CV and click the Update cvtermpath button.
Now, enable the Tripal GO module. Open a new tab and navigate to Administer → Site Building → Modules. Enable the Tripal GO module.
Now go to the feature page and see the list of GO terms associated to that feature.
Blast Module
First, enable the Blast module navigate to Administer → Site Building → Modules and enable the Tripal Blast module.
Interpro analysis for feature ‘FBgn0040037’. Download these results from this link and unzip:
cd /home/gmod/Downloads
unzip JYalpha.fasta_vs_sprot.zip
Next, we need to create a new analysis for the SwissProt results. Do this by clicking Create Content on the left sidebar and then click Analysis: Blast. In the form that appears provide the following details:
- Analysis Name: Blastx FBgn0040037 vs Swissprot
- Program: Blastx
- Program Version: 2.2.18
- Time Executed: Today
- Database: UniProt/SwissProt
- Blast xml File: /home/gmod/Downloads/JYalpha.fasta_vs_sprot.xml
- Check: Submit a job to parse the xml output into analysisfeatureprop table
- Parameters: ‘blastall -p blastx -d uniprot_sprot.fasta -i JYalpha.fasta -e 1e-6 -m 7 -o JYalpha.fasta_vs_sprot.xml’
Save the Analysis and run the new job:
cd /var/www; sudo php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Now return to the feature: FBgn0040037 and see the results.
However, the blast doesn’t look quite the way we want. We want to fine tune it. Navigate to Administer → Tripal Management → Analysis. Scroll down to the Blast Parser Settings.
Select the Uniprot/SwissProt database and then set the regular expressions accordingly:
| Title: | ExPASy Swissprot |
| Hit Name: | .*?\(.*?)\s.*?$ |
| Hit Description: | .*?\.*?\s(.*)$ |
| Hit Accession: | (.*?)\.*?\s.*?$ |
Now return to the feature page and view the blast results. They look better. But we want to make the results link out. Navigate to Administer → Tripal Management → DB. On the 6th page, click edit next to Uniprot/SwissProt and add the following:
- URL: http://us.expasy.org/
- URL prefix: http://us.expasy.org/cgi-bin/niceprot.pl?
Now return to the feature page and view the blast results. The match names now link to the external database.
InterPro Analysis
First, enable the Interpro module navigate to Admminister → Site Building → Modules and enable the Tripal Interpro module.
Interpro analysis for feature ‘FBgn0040037’. Download these results from this link and unzip:
cd /home/gmod/Downloads
unzip JYalpha.fasta.ipr.zip
Next, we need to create a new analysis. Do this by clicking Create Content on the left sidebar and then click Analysis: Interpro. In the form that appears provide the following details:
- Analysis Name: InterProScan Analysis of FBgn0040037
- Program: InterProScan
- Program Version: 4.5
- Time Executed: Today
- Interpro Output File: /home/gmod/Downloads/JYalpha.fasta.ipr.html
- Check: Submit a job to parse the interpro HTML output
- Check: Load GO terms to the database
- Parameters: ‘iprscan -cli -i JYalpha.fasta -o JYalpha.fasta.ipr.html -format html -goterms -ipr’
Save the Analysis and run the new job:
cd /var/www; sudo php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Now return to the feature: FBgn0040037 and see the results.
The Intepro analysis has GO terms associated with it. Therefore the GO module can visualize those. One additional feature of the GO module is that it can provide a summary of GO annotations for a the organism page. Go to the organism page. There is box titled Gene Ontology Summary but it has no information. This information requires a materialized view. We need to update the view. Navigate to Administer → Tripal Management → MViews. There are three materialized views added by the CV and GO modules that need updating: ‘cv_root_mview’, ‘go_count_analysis’, ‘go_count_organism’. Now manually run the job:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Now, return to the organism page and scroll to the bottom. There now appears a drop down box where the analysis can be selected. Once select pie charts appear showing the report. Open the tree and click any of the go terms in the tree.
Click the Show on Menu link at the bottom of the GO report. The report now appears as a secondary menu link on the top of the page. It can be returned to the bottom of the main page by clicking the Remove from menu link.
Finally, click on the Analysis link at the top of the page. The analysis now appears with details.
Searching
We have a search box at the top of the page but we would like to add a menu item for searching. Navigate to the Administer → Site building → Menus → Primary Links page and click the Add Item near the top of the page.
Enter the following into the fields
- Path: search/node
- Menu link title: Search
A new Search tab appears in the main menu.
Check the status of the search indexing by navigating to Administer → Site Configuration → Search Settings. For searching to work correctly the site must be 100% indexed. If not let’s do that now by manually running the Drupal cron:
/usr/bin/wget -O - -q http://localhost/cron.php
Additionally, in the blast results just added we want to have the term ‘catalytic activity’ to be searchable, but if we search on that we may get zero results. We would like for both terms to be searchable. To do this, we need to reindex the features.
How to reindex all content
Navigate to Administer &rar; Tripal Management → Features and click the Reindex all feature nodes button. This will add a Job to Tripal.
Rather than wait for the Tripal Job cron to launch we will do it manually:
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php fly_admin
Check the status of this job on the Jobs page.
We should now be able to search for ‘catalytic activity’.
Anytime major changes are made to the content on feature pages the feature should be re-indexed.
Advanced Searching
Drupal uses “Taxonomy” to provide filtering of search results. In the case of this demo this level of advanced searching wouldn’t make sense because we only have one organism and few feature types but we’ll walk through how to do it.
An example of advanced searching with multiple taxonomy terms: http://www.fagaceae.org/search
- First, navigate to the Search page and open the advanced options. It provides some default filtering mechanisms.
-
Navigate to Administer → Tripal Management → Features. Scroll down to the Set Taxonomy section and check the boxes Organism Name and Feature Type and click the button Set/Reset Taxonomy for all feature nodes. This will create a Tripal job. Let’s launch the job manually rather than wait for the cron to kick in.
cd /var/www; php ./sites/all/modules/tripal_core/tripal_launch_jobs.php yeast_admin
- Check the status of the job on the Tripal Jobs page. When 100% complete return to the Search page and refresh. You’ll notice now a box for filtering search results by organism and by feature type. Search for the Feature ‘YAL061W’. click on the result and scroll to the bottom of the page. You’ll notice some “tags” have been added to the page. These are the Drupal taxonomy tags that have been assigned to the feature page.
- Navigate to Administer → Content Management → Taxonomy. You’ll see the new vocabularies and terms.
Libraries & other Analysis
We do not have libraries for this example but they function in much the same way.
Subsets of features can be synced on the analysis and library administration pages when those modules are installed and features are associated in chado.
Incorporate JBrowse
Exercise:
- Create a new page titled ‘JBrowse’
- Add the following in the body:
<iframe id="gbrowseFrame" width="100%" height="1500px" frameborder="0" src="/jbrowse/"> - Set the Input type to Full HTML.
- Save the page.
- Create a menu item
This JBrowse won’t contain to the sequence in our demo database but for purposes of demonstration this is adequate.
Example Customizing Content
In the case of the organism template, the organism abbreviation is not
present. This can easily be added by editing the
node-chado_organism.tpl.php template file.
cd /var/www/sites/all/themes/theme_tripal
Add this table row to the code:
<tr>
<th>Abbreviation</th>
<td><?php print $node->abbreviation?></td>
</tr>
With PHP, CSS and JavaScript abilities the look-and-feel of chado content can be completely changed.
Writing your own Module
Not yet in the User’s Guide
Drupal API
Concepts
- The Tripal Core provides all necessary functionality to intergate with Tripal.
- Drupal API provide the rest
Anatomy of a Drupal module
- module name (directory name)
- a .info file describing the module
- a .install file the gets executed when the modules if first installed and also when uninstalled.
- a .module file the code for the module.
- templates: the look-and-feel should be separated from the code if possible.
Example .info file
tripal_feature.info
; $Id: tripal_feature.info,v 1.4 2009/10/01 17:52:24 ccheng Exp $
name = Tripal Chado Feature
description = A module for interfacing the GMOD chado database with Drupal, providing viewing, inserting and editing of chado features.
core = 6.x
project = tripal_feature
package = Tripal
dependencies[] = tripal_core
dependencies[] = tripal_organism
dependencies[] = search
dependencies[] = path
dependencies[] = tripal_cv
version = "6.x-0.2b-m0.2"
Example .install file
tripal_feature.install
Drupal hooks: {node}_{hook_name} or {module}_{hook_name}
The important functions:
- function tripal_feature_install(): actions to do on install
- function tripal_feature_schema(): returns the database schema so that Drupal knows what it is
- function tripal_feature_uninstall(): actions for cleanup when the module is uninstalled
See the code here: /var/www/sites/all/modules/tripal_feature
Example .module file
tripal_feature.module
The important module-centric functions:
- function tripal_feature_admin(): generates the administration form for the module.
- function tripal_feature_admin_validate($form, &$form_state): validates the admin form input.
- function tripal_feature_node_info(): defines new nodes for Drupal (in this case chado_feature).
- function tripal_feature_perm(): defines new permissions for the module.
- function tripal_feature_menu(): defines new menu items
- function tripal_feature_theme(): when theming must be handled by the module (e.g. search results).
- function tripal_feature_nodeapi(&$node, $op, $teaser, $page): used to place content on other nodes pages.
The important node-centric functions:
- function chado_feature_access($op, $node, $account): permission checking for the nodes
- function chado_feature_insert($node): code to execute on insert of a new node
- function chado_feature_delete($node): code to execute on delete of a node
- function chado_feature_update($node): code to execute on update of a new node
- function chado_feature_form ($node,$param): generates the form when editing or adding a new node
- function chado_feature_validate($node): validate the form input on insert or update.
- function chado_feature_load($node): loads the node data object when viewed
- function chado_feature_view ($node, $teaser = FALSE, $page = FALSE): set’s in-module theming.
Tripal API
Tripal provides the following functions for integrating with Tripal:
tripal_create_moddir($module_name)
Every tripal module is expected to have it’s own data directory. This directory gets created in the ./sites/default/files/tripal directory. This function creates that directory.
tripal_get_moddir($module_name)
Call this function to get the path for the directory that was created above.
tripal_add_job ($job_name,$modulename,$callback,$arguments,$uid, $priority = 10)
This function adds a new job is added to Tripal. An explanation of the argument is as follows:
- job_name: The name of the job as it appears to the user
- modulename: The name of the Tripal module submitting the job (e.g. tripal_feature, tripal_organism, etc).
- callback: The name of a function in the caller’s module that should be called when the job is executed.
- arguments: An array of arguments to pass to the callback function when executed.
- uid: the user id of the user executing the job.
- priority: sets the priority of the job. The lower the number the higher the priority. Administrative jobs should always be 10 or less. User submitted jobs should always be above 10.
Note: This function will always pass in the job id as the first argument to the callback function. Therefore all callback functions should have a jobid argument first.
tripal_job_set_progress($job_id,$percentage)
Allows the callback function for the job to set the progress of the job. The first argument is the job id that get’s passed into the callback by Tripal and the percentage is a value between 0 and 100.
tripal_get_module_active_jobs ($modulename)
Allows a module to see if it has any active jobs currently executing. A list of jobs is returned.
tripal_add_cvterms ($name,$definition,$cv_name = ‘tripal’,$db_name=’tripal’
Adds a CVterm to the chado database. Terms added using this function are added by default to the ‘tripal’ CV and associated with the ‘tripal’ database in Chado. This function allows modules to create cvterms behind-the-scenes to support the data management they provide. This function is particularly useful in _install hooks of .install files.
function tripal_add_mview ($name,$modulename,$mv_table,$mv_specs,$indexed,$query,$special_index)
Programatically adds a materialized view to chado. This view can then be used behind-the-scenes by the module to help speed data queries. This function is particularly useful in _install hooks of .install files. An explanation of the argument is as follows
- name: The name of the materialized view.
- modulename: The name of the module submitting the materailzed view (e.g. ‘tripal_library’)
- mv_table: The name of the table to add to chado. This is the table that can be queried.
- mv_specs: The table definition
- indexed: The columns that are to be indexed
- query: The SQL query that loads the materialized view with data
- special_index:
Note: These views are managed by Tripal and only the view itself is stored in Chado. This is different from the materialized views that comes with chado 1.0, although the outcome is the same. Any materialized views created with Tripal are not compatible with the Chado scripts for updating materialized views.
Adding a new analysis method
Not yet fully defined, however, the tripal_analysis module will provide a set of “core” functions for all analysis modules. Documentation and finalized analysis API coming soon.
Exercise
Setup
Step 1
Add a publication to Chado
Let’s create a very simple module that will place publications on a feature page if an association exists in chado. We won’t be creating a data entry method so we’ll have to manually add an example publication. Using phpPgAdmin, execute the following SQL statements (together as a block):
Alternatively, log on using the command-line:
psql -U fly_admin -d chado_dmel
Now execute the following SQL statements
INSERT INTO cv (NAME) VALUES ('pub_demo');
INSERT INTO db (NAME) VALUES ('pub_demo');
INSERT INTO dbxref (db_id,accession,version,description)
VALUES ((SELECT db_id FROM db WHERE NAME = 'pub_demo'),
'pub0001',' ','demo publication accession');
INSERT INTO cvterm (cv_id,name,dbxref_id)
VALUES ((SELECT cv_id FROM cv WHERE NAME = 'pub_demo'),
'journal',
(SELECT dbxref_id FROM dbxref WHERE accession = 'pub0001'));
INSERT INTO pub (title,volumetitle,volume,issue,pyear,pages,uniquename,type_id)
VALUES ('Multiple SET methyltransferases are required to maintain normal heterochromatin domains in the genome of Drosophila melanogaster',
'Genetics','181','4','2009','1303-1319','demo_pub',
(SELECT cvterm_id FROM cvterm CVT INNER JOIN CV ON CVT.cv_id = CV.cv_id
WHERE CV.name = 'pub_demo' AND CVT.name = 'journal'));
INSERT INTO pubauthor (pub_id,rank,surname,givennames)
VALUES ((SELECT pub_id FROM pub WHERE uniquename = 'demo_pub'),0,'Brower-Toland','et al.');
INSERT INTO feature_pub (feature_id,pub_id)
VALUES ((SELECT feature_id FROM feature WHERE uniquename = 'FBgn0040037'),
(SELECT pub_id FROM pub WHERE uniquename = 'demo_pub'));
These statements do the following:
- Adds a db and cv. It’s not real, but suits our purposes for the example.
- Adds a publication and author
- Associates the publication with the feature YAL061W.
Step 2
create the module directory
Change directories to our Drupal install directory where we installed the Tripal modules:
cd /var/www/sites/all/modules
Create a directory for our module. The directory name should be identical to our module name.
mkdir tripal_pubs
cd tripal_pubs
Create tripal_pubs.info file
The first step to creating a module is to define our .info file that provides information to Drupal about our module.
Design considerations:
- Our module is dependent on the Tripal core (all modules are) and the Tripal feature (we want to tie our pubs to features).
- We want to add this module to the Tripal package.
Create a tripal_pubs.info file with the following content:
; $Id:
name = Tripal Pubs
description = A module for displaying publications on feature pages
core = 6.x
project = tripal_pubs
package = Tripal
version = 6.x-0.2-m0.1
dependencies[] = tripal_core
dependencies[] = tripal_feature
Create tripal_pubs.install file
We do not have any functionality that needs to be performed when the module is installed, but we’ll create a shell with proper drupal hooks so that in the future if we do want to add code to these modules we can.
<?php
//$Id:
/*******************************************************************************
* Implementation of hook_install().
*/
function tripal_pubs_install() {
}
/*******************************************************************************
* Implementation of hook_uninstall().
*/
function tripal_pubs_uninstall() {
}
/*******************************************************************************
* Implementation of hook_schema().
*/
function tripal_pubs_schema() {
$schema = array();
return $schema;
}
Create the tripal_pubs.module file
Our module will be simple. It will add content to a feature page if there are publications associated with the feature. However, we’ll add many important hooks for demonstration purposes.
Cut and paste the following code into a new tripal_pubs.module file.
We’ll discuss each function…
<?php
/*******************************************************************************
* Implementation of hook_init()
*/
function tripal_pubs_init() {
}
/*******************************************************************************
* Implementation of hook_perm()
*/
function tripal_pubs_perm(){
return array(
'access chado_publication content',
'create chado_publication content',
'delete chado_publication content',
'edit chado_publication content',
);
}
/*******************************************************************************
* Implementation of hook_node_info()
*/
function tripal_pubs_node_info() {
$nodes = array();
$nodes['chado_publication'] = array(
'name' => t('Publications'),
'module' => 'chado_publication',
'description' => t('A publication from the chado database'),
'has_title' => FALSE,
'title_label' => t('Publication'),
'has_body' => FALSE,
'locked' => TRUE
);
return $nodes;
}
/*******************************************************************************
* Implementation of hook_block()
*/
function tripal_pubs_block($op = 'list', $delta = '0', $edit = array()){
switch($op){
case 'list':
$blocks[0]['info'] = t('Publications');
return $blocks;
case 'view':
if(user_access('access chado_publication content')){
$items[] = t("This block is not yet setup.");
$block['subject'] = t('Publications');
//We theme our array of links as an unordered list
$block['content'] = theme('item_list', $items);
}
return $block;
}
}
/*******************************************************************************
* Implementation of hook_menu()
*/
function tripal_pubs_menu() {
$items = array();
$items['publications'] = array(
'menu_name' => ('primary-links'), //Enable the 'Publications' primary link
'title' => t('Publications'),
'page callback' => 'tripal_pubs_page',
'access arguments' => array('access chado_publication content'),
'type' => MENU_NORMAL_ITEM
);
// the administative settings menu
$items['admin/tripal/tripal_publication'] = array(
'title' => 'Publications',
'description' => 'Manage integration of Chado publications including associated features',
'page callback' => 'drupal_get_form',
'page arguments' => array('tripal_pubs_admin'),
'access arguments' => array('administer site configuration'),
'type' => MENU_NORMAL_ITEM,
);
return $items;
}
/*******************************************************************************
* Implementation of hook_admin()
*/
function tripal_pubs_admin () {
// provide a form for administrative settings
$form = array();
return system_settings_form($form);
}
/*******************************************************************************
* Implementation of hook_admin_validate()
*/
function tripal_pubs_admin_validate($form, &$form_state) {
// validate the admin form submission
}
/*******************************************************************************
* Implementation of hook_insert()
*/
function chado_publication_insert($node){
// add publication to chado database
}
/*******************************************************************************
* Implementation of hook_delete()
*/
function chado_publication_delete($node){
// remove publication from chado database
}
/*******************************************************************************
* Implementation of hook_update()
*/
function chado_publication_update($node){
// update a publication record in the chado database
}
/*******************************************************************************
* Implementation of hook_access()
*/
function chado_publication_access($op, $node, $account){
if ($op == 'create') {
return user_access('create chado_publication content', $account);
}
if ($op == 'update') {
if (user_access('edit chado_publication content', $account)) {
return TRUE;
}
}
if ($op == 'delete') {
if (user_access('delete chado_publication content', $account)) {
return TRUE;
}
}
if ($op == 'view') {
if (user_access('access chado_publication content', $account)) {
return TRUE;
}
}
return FALSE;
}
/*******************************************************************************
* Implementation of hook_form()
*/
function chado_publication_form ($node, $param){
$form = array();
$form['title']= array(
'#type' => 'textfield',
'#title' => t('Publication Title'),
'#required' => TRUE,
'#default_value' => ' ',
'#description' => t('Enter the title for the publication'),
'#weight' => 1,
'#maxlength' => 255
);
// add additional fields to the form that are needed to populate the
// chado tables
return $form;
}
/*******************************************************************************
* Implementation of hook_load()
*/
function chado_publication_load($node){
$previous_db = db_set_active('chado'); // use chado database
$sql = "SELECT * ".
"FROM pub P ";
$pubs = db_fetch_object(db_query($sql));
db_set_active($previous_db); // now use drupal database
$additions->pubs = $pubs;
// we would additionally want to pull out the authors for the publication
// as well
}
/*******************************************************************************
* Implementation of hook_view()
*/
function chado_publication_view ($node, $teaser = FALSE, $page = FALSE) {
// this function provides instructions for establishing the way the
// publication page will look.
}
/*******************************************************************************
* Callback for the 'publications' menu item
*/
function tripal_pubs_page(){
return 'This is where custom content would go if we needed it.';
}
/*******************************************************************************
* Implementation of hook_nodeapi()
*/
function tripal_pubs_nodeapi(&$node, $op, $teaser, $page) {
switch ($op) {
case 'view':
// Abort if this node is not one of the types we should show.
if (strcmp($node->type,'chado_feature') != 0) {
break;
}
// Add pubs to the content item if it's not a teaser
if (!$teaser && $node->feature->feature_id) {
$node->content['tripal_pubs_form'] = array(
'#value' => theme('tripal_pubs_feature_pub_list', $node),
'#weight' => 4
);
}
}
}
/*******************************************************************************
* Implementation of hook_theme()
*/
function tripal_pubs_theme () {
return array(
'tripal_pubs_feature_pub_list' => array (
'arguments' => array('node'),
)
);
}
/*******************************************************************************
* The theme function that sets the content to be added
*/
function theme_tripal_pubs_feature_pub_list ($node) {
$feature = $node->feature;
// get the list of publications that are associated with this
// feature
$sql = "SELECT * ".
"FROM pub P ".
" INNER JOIN feature_pub FP ON FP.pub_id = P.pub_id ".
"WHERE FP.feature_id = %d";
$previous_db = db_set_active('chado'); // use chado database
$pubs = db_query($sql,$feature->feature_id);
db_set_active($previous_db); // now use drupal database
// generate the content that will appear on the feature page
// first we'll make the normal Tripal expandable box for this
// content to go into
$content = "".
"".
"<h3>Publications</h3>".
"feature_id\">";
// iterate through the publications, get the authors and print
// the references
while($pub = db_fetch_object($pubs)){
$sql = "SELECT * FROM pubauthor WHERE pub_id = %d";
$previous_db = db_set_active('chado'); // use chado database
$authors = db_query($sql,$pub->pub_id);
db_set_active($previous_db); // now use drupal database
while ($author = db_fetch_object($authors)){
$content .= "$author->surname, $author->givennames; ";
}
$content .= "$pub->title. <em>$pub->volumetitle</em> $pub->pyear $pub->volume($pub->issue):$pub->pages<br>";
}
$content .= "<br>";
return $content;
}
Note… fix the < div> and < h3>tags in the function immediately above and remove the spaces. Mediawiki was giving problems with these so they should be fixed.
Facts about “Tripal Tutorial 2010”
| Has topic | Tripal + |