JBrowse
From GMOD
JBrowse is a genome browser with almost entirely AJAX-based interfaces. JBrowse renders most tracks using client side JavaScript and JSON as its data transfer format. JBrowse is currently under development by the Ian Holmes Lab at UC Berkeley. It is expected to have a production release in 2009. Most development has been done by Mitch Skinner. JBrowse was formerly known as GBrowse 3. However, this name was changed to JBrowse to avoid confusion: JBrowse is an alternative to GBrowse, not a successor.
Requirements
- What is needed?
Documentation
Presentations
- Image:GBrowse3GMOD2008.pdf - Presentation by Ian Holmes at July 2008 GMOD Meeting. See the talk's summary for more.
- JBrowse - Presentation by Mitch Skinner at the January 2009 GMOD Meeting. See the talk's summary for more.
Downloads
Demo & Screenshots
See the demo site for a bleeding edge demo.
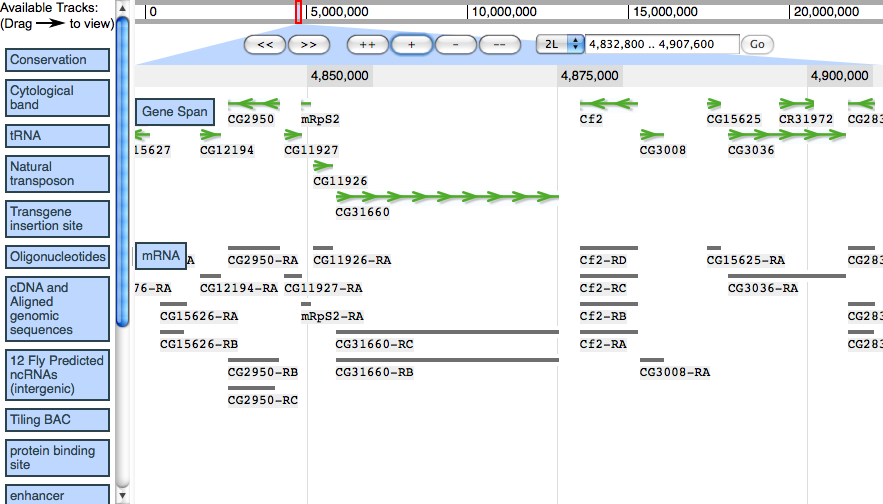 Demo screenshot from 2009/02/20, showing region of Drosophila melanogaster 2L.
Demo screenshot from 2009/02/20, showing region of Drosophila melanogaster 2L.
Mailing List
- gmod-ajax@lists.sourceforge.net: Discussion of development, installation problems, etc.
- Subscribe to gmod-ajax
- gmod-ajax archives
Contact
Facts about "JBrowse"RDF feed
| Available on platform | web + |
| Has URL | http://jbrowse.org/install/ +, http://jbrowse.org +, http://twitter.com/usejbrowse +, http://github.com/GMOD/jbrowse +, http://jbrowse.org/demos +, http://icemangenome.net/%E2%80%8E +, http://genomesunzipped.org/jbrowse +, http://beetlebase.org + and http://www.medicinalgenomics.com/the-jane-ome/ + |
| Has description | Browse the genome of Ötzi the ice man + and JBrowse is a genome browser with a fully d … JBrowse is a genome browser with a fully dynamic AJAX interface, being developed as the eventual successor to GBrowse. It is very fast and scales well to large datasets. JBrowse is javascript-based and does almost all of its work directly in the user's web browser, with minimal requirements for the server.
Features[edit]
|
| Has development status | active + |
| Has input format | GFF3 +, BED +, FASTA +, WIG +, BedGraph +, Bio::DB::* +, UCSC +, Wiggle +, BigWig + and BAM + |
| Has licence | LGPL + and Artistic License 2.0 + |
| Has logo | JBrowseLogo.png + |
| Has software maturity status | mature + |
| Has support status | active + |
| Has title | JBrowse demos +, Ice Man Genome +, Genomes Unzipped: Public Personal Genomics +, BeetleBase + and The Jane-Ome, medicinal marijuana project + |
| Has topic | JBrowse + |
| Is open source | Yes + |
| Link type | download +, social media +, website +, source code +, demo server + and wild URL + |
| Release date | 2008 + |
| Tool functionality or classification | Genome visualization + |
| Written in language | Javascript + and Perl + |
| Has subobjectThis property is a special property in this wiki. | JBrowse#http://jbrowse.org/install/ +, JBrowse#http://jbrowse.org +, JBrowse#http://twitter.com/usejbrowse +, JBrowse#http://github.com/GMOD/jbrowse +, JBrowse#http://jbrowse.org/demos +, JBrowse#http://icemangenome.net/ +, JBrowse#http://genomesunzipped.org/jbrowse +, JBrowse#http://beetlebase.org + and JBrowse#http://www.medicinalgenomics.com/the-jane-ome/ + |