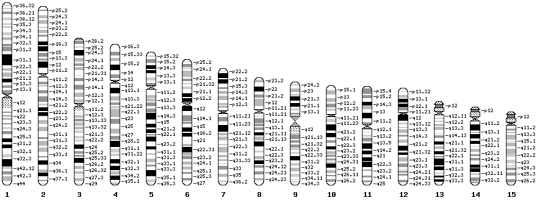
GBrowse karyotype
From GMOD
GBrowse Karyotype documentation and demo installation
Note on installation
- GBrowse_karyotype is not part of the GBrowse distribution
- It should be checked out from Subversion, or downloaded from here.
- Install as you would GBrowse, after you have installed GBrowse 1.69, or 1.68, or a SVN checkout of the 'stable' branch
cd GBrowse_karyotype perl MakeFile.PL make install
Cytoband Data
Follow this link for an example of a Perl script that will download and format cytband data from EnsEMBL